What is ENCODE ?
ENCODE, an acronym for Encyclopedia of DNA Elements, is a publicly funded research project
initiated in 2003 by National Human Genome Research Institute (NHGRI). Results of the pilot phase was published in June 2007 in Nature.
After the success of pilot phase, ENCODE was given a go ahead to scale up and analyze the entire genome.
It has extended our learnings from work completed in 2001 by the Human Genome Project to sequence all 3 billion chemical base pairs in human DNA.

Image Credit : www.genome.gov
In the 1st week of September, 2012, the results of the ENCODE project were published in a bouquet of 30 papers across multiple journals. The project was propelled by a collaboration of about 450 scientists across 32 labs from around the world.
Key scientific insights and implications
ENOCDE has essentially managed to map a dashboard of genetic switches which are involved in switching on and off genes coding for proteins. Nearly 4 million genetic switches are present in the genetic regulatory system. Findings of the project indicate that some of these switches may have a role in regulation of biochemical functions and/or involved in diseases.
Several diseases may be linked to a bunch of genetic switches, which means drug companies may be able to find better cures for cancer by focusing on relatively few targets.
State of science writing
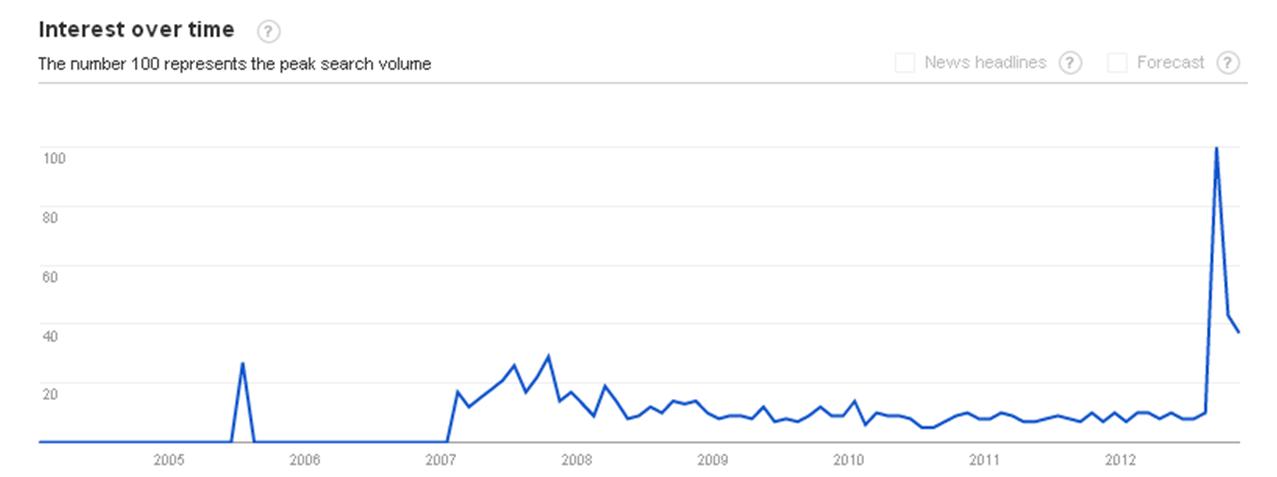
Image Credit : Google Trends. Searching "Project ENCODE" in Google Trends confirms the massive publicity it received after the papers were published in various journals in the 1st week of Septemeber, 2012.
There have been major concerns on confusing usage of terminologies and misinterpretation of findings of ENCODE by some sections of the media. Experts have called for serious introspection of current state of science writing and major overhauling of the same in near future.
Big data, Big Collaboration
Implementation of ENCODE was enabled by the recent growth in big data processing and DNA sequencing technology. Scientists in the U.K., U.S, Spain, Singapore and Japan generated and analysed over 15 terabytes of raw data, different chunks of which are stored in massive data processing centres.
Such projects have highlighted the need for addressing the computing needs of small research groups which are not well funded. In order to solve this problem, EU has started the CHAIN project through which any researcher would be able to access a global computing grid.
New way of publishing research papers and data
ENCODE papers published were made available via a nice graphical user interface on the web and as an iPad app. The papers were clubbed into various threads like "Characterization of network topology", "Transcription factor motifs" etc. for easy navigation across different topics.
Big and small research projects
Increasing squeeze on the availability of funding for research has led to debates on the scientific and economic value of big research projects as compared to smaller projects. Concerns have been raised about the possibility of lesser access to funds for smaller research projects due to inclination of grant agencies to fund big budget projects. Huge number of scientific discoveries made by individual investigators also makes a strong case for small research projects. One would expect the debate to continue and ultimately lead to betterment of science, and good use of public funds. No wonder measuring research impact is a hot field of study these days as evident from growing number of scholarly metrics.
Government funded and private funded research
Some analysts have pointed out the need for funding basic research projects by governments as big budget and long term projects like ENCODE are difficult to sustain financially in a corporate environment. In spite of financial challenges, companies are exploring innovative business models and forming consortium to facilitate basic research.
Future Outlook
Making sense of enormous amount of data produced by ENCODE will take time. Better understanding of data is expected to spawn new research directions leading to better understanding of human genome. It may help in the reversal of declining pharma R&D productivity and lead to advancement of personalized medicine. Next phase of ENCODE will carry out deeper exploration of human genome to answer the unanswered questions so far. Meeting the expectations arising out of the recent publications would be a major challenge for this endeavour.
References
-----------------
1. http://www.genome.gov/10005107
2. http://www.nature.com/encode/
3. http://www.huffingtonpost.com/2012/09/05/human-genome-encode-dna-genes_n_1858281.html
4. http://www.huffingtonpost.com/athena-andreadis-phd/junk-dna-junky-pr_b_1900243.html
5. https://itunes.apple.com/us/app/nature-encode/id553487333?mt=8
7. http://www.isgtw.org/feature/picking-long-tail-scientific-computations
8. http://www.rsc.org/chemistryworld/2012/10/h-index-research-metric-prediction
9. http://www.michaeleisen.org/blog/?p=1179
10. http://www.genomeweb.com/encode-expands-30m-new-projects-data-centers
11. http://www.nature.com/nrd/journal/v11/n3/full/nrd3681.html




